The body plan of multicellular organisms is under genetic control.
Complete the passage below using the most appropriate words from the list.
analogous archaea development DNA domains
homeobox homologous homozygous kingdoms operon
phyla plant preserved prokaryotes regulator
ribosomes transcription translation
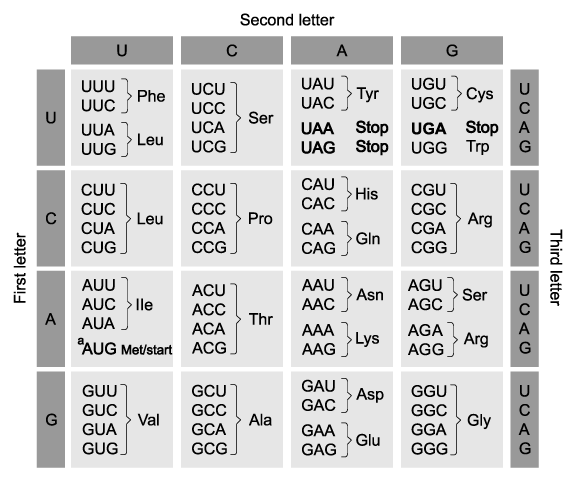
The development of body plan in eukaryotic organisms is controlled by ................................. genes. These genes code for proteins that are able to bind to ................................... and turn specific genes on and off and are known as ......................................... factors. These proteins contain a sequence of base pairs that varies little between species within the animal, ............................................. or fungus ..........................................
Investigations into the activity of genes that control body plan frequently use fruit flies and mice.
One reason fruit flies are used is that there are fewer public concerns about the ethics of using flies.
(i) Suggest two other reasons why fruit flies are chosen for research into genes controlling the development of body plan.
[2]
(ii) There are some public concerns about the ethics of using mice in these investigations.
Suggest two reasons why mice are chosen as a suitable species for investigation.
[2]
Was this exam question helpful?